Agrigenomics Yields a Next-Gen Cornucopia

Benefits of genome editing and molecule-sensing technologies in agriculture include more sustainable crops and healthier foods.
Consumers may soon begin purchasing fun-sized fruits and vegetables, as well as processed foods that incorporate healthier ingredients, such as oils that are relatively free of “unhealthy” fats. And producers may be able to grow crops that are drought- and flood-tolerant, yield more per acre, and are easier to harvest and transport—and are tastier, more nutritious, and less allergenic, too. These are just a few of the possibilities that are being realized thanks to recent applications of gene editing technology in crop science.
Gene editing technology takes agricultural biotechnology beyond transgenic technology, which transfers “as is” genes from one species to another. Essentially, gene editing is more refined than transgenesis. This difference may justify the view that gene edited plants are not, like transgenic plants, genetically modified organisms (GMOs). Regulatory requirements and consumer attitudes may hang in the balance.
Not only has transgenesis been slapped with the GMO label, it has proven to be a time-consuming and expensive approach. It has been applied to just a handful of row crops, that is, crops representing large markets targeted by large multinationals undaunted by large commitments—10-year development times and $100-million-plus investments.
Now that gene editing tools are available to crop scientists, the engineering of improved plants is becoming faster and easier. For example, CRISPR tools are accelerating development because they are economical and provide advanced capabilities such as multiplexing. In fact, gene editing is democratizing the development of engineered plants. Not only is the technology being adopted by large, established players such as Syngenta, Bayer, BASF, and Corteva, it is driving the emergence of small companies such as Calyxt and Pairwise Plants. Other emerging industry players, like Ontera, are developing robust tools that enable the elucidation of disease and resistance mechanisms and the fast and precise molecular identification of therapeutic targets.
But CRISPR isn’t a magic wand that makes visions come true. There are obstacles it doesn’t bypass. Random genomic mutations, for example, occur naturally during every typical breeding cycle. And gene editing can introduce off-target effects, where the editing machinery performs incorrect or incorrectly located edits.
Some of these difficulties can be eliminated through line selection or adverse selection. These correctives, however, can be time consuming and expensive. Consequently, off-target effects could turn the gene editing vision into an unreachable dream. If the vision is to be realized, plant scientists will need to cultivate “out of the box” thinking. The main challenges are increasing gene editing’s efficiency and specificity and improving the plant-regeneration process.
Optimizing the process
Gene editing reagents are often still delivered to plant callus cells in culture using decades-old technology, such as Agrobacterium- and biolistics (transformation via ballistic DNA delivery)-based approaches. The edited callus cells are then allowed to proliferate, and hormones are applied to eventually induce the formation of a shoot, then a root, and finally (maybe six to nine months later) a plant.
Each step—delivery, editing, and regeneration—is inefficient. Regenerating a plant from cells in culture is a massive bottleneck and must be optimized for each species. Even within a species, different strains or varieties may regenerate well, while others, such as elite varieties, may be recalcitrant.
“Methods that obviate the need for tissue culture would be a big boon,” says Daniel Voytas, PhD, professor of genetics, cell biology, and development at the University of Minnesota; director of the Center for Precision Plant Genomics; and co-founder and chief scientific officer at Calyxt. “One approach is to sidestep culture altogether.”
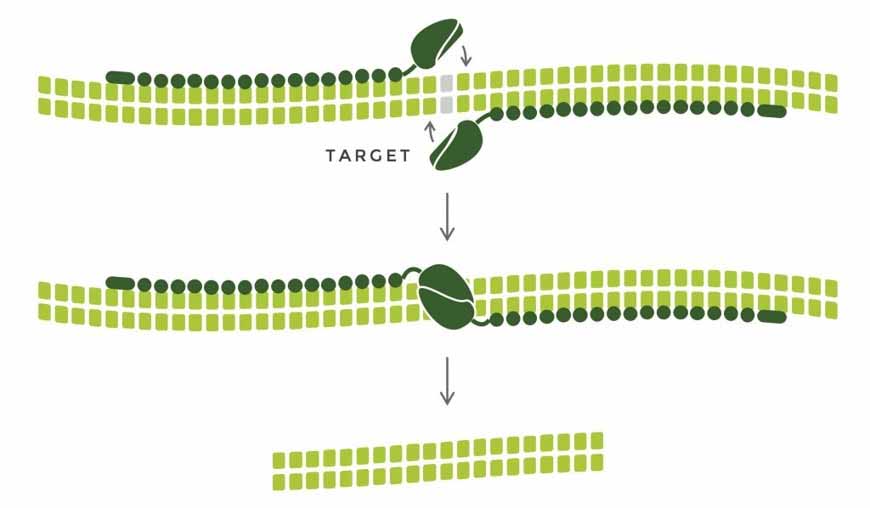
Voytas and his research team at the University of Minnesota work on optimizing gene editing efficiency. “We prune a plant and add gene editing reagents to the pruned sites along with developmental regulators to induce shoot formation,” he explains. “In some cases, this works, eliminating the need to use sterile technique on single cells, to identify the one cell that has been edited, and to regenerate a plant, which takes months.”
The Voytas laboratory is also experimenting with DNA and RNA viral vectors to infect germline cells. Since plants have mechanisms to eliminate viruses, there is typically no residual infection when the regenerated plant produces seed. Regardless, depending on the delivery approach, careful screening of the progeny ensures the absence of transgenes or virus.
“My colleagues are using gene editing to answer a wide variety of questions about plant biology and to develop new crop varieties, including niche crops such as berries, fruits, and vegetables,” notes Voytas. “You are going to see dozens or hundreds of varieties coming from academic laboratories. You never saw an academic laboratory work on a GMO crop.”
At vertically integrated Calyxt, the focus is on creating healthier food ingredients. Calyxt’s first commercially available product is Calyno high-oleic soybean oil. The company creates new varieties using gene editing, contracts with farmers to grow the crop, then buys back the grain and processes it for sale as a food ingredient, sharing profits with the growers. The company’s next focus is on wheat; other crops, such as alfalfa, are developed with partners.
Fruits and vegetables
“Diet is a major contributor to many of the health problems we face as a society,” says Tom Adams, PhD, chief executive officer, Pairwise Plants. “Two billion people have diseases or prediseases related to unhealthy diets. If we all ate one more serving of fruit a day, heart disease deaths would decrease by one in seven.
“Our mission is to use gene editing technology to increase fresh produce consumption in America, and eventually worldwide, by removing barriers [to the improvement of] taste, quality, availability, and convenience.”
Of course, humans have been using artificial selection on plants to remove just these barriers for 12,000 years in a process known as agriculture. What’s different now is the range of possibilities resulting from the new genetic techniques. “We have been limited to the phenotypes that arise in nature or from breeding programs,” Adams explains. “Now, with gene editing, we can maximize phenotypes for productivity and health, especially in clonally reproduced and niche crops.”
Edited versions of berries and cherries, however, have a different commercialization pathway than that of row crops. In a row crop, multiple generations are made and their phenotypes evaluated in a breeding program. Consumer crops in contrast are clonally propagated. Gene editing makes it much easier to introduce a trait without all the traditional backcrosses.
Adams adds, “CRISPR technology is inexpensive and readily available, and although there are still some hurdles to get it to work in all crops, it allows business models that focus on smaller markets and not just globally traded grains.”
Knockouts are relatively easy to make, but it is more difficult to make fine-tuned changes in sequences. Pairwise has licensed rights to a base editing technology, a modification of CRISPR, developed by David R. Lui, PhD, a professor of chemistry and chemical biology at Harvard University and one of Pairwise’s co-founders. For base editing, the Cas9 enzyme, the part of CRISPR technology responsible for cutting the DNA double helix, is inactivated and replaced with a chemical modifier to make fine-tuned C-to-T or A-to-G changes at high efficiency.
Currently in the United States, gene editing does not fall under the same strict food regulations as GMO products; however, the absence of transgenes in the final product and the similarity of the product to a breeding result must be demonstrated. Globally, many countries are following a similar regulatory path. Notable exceptions include countries in Europe.
Row crops
Transformation of elite varieties, those with a preferred set of traits, tends to be troublesome because many of these varieties do not regenerate in culture. To circumvent the regeneration problem, laboratories will transform a relatively amenable laboratory variety. If an edit of interest is generated with the desired trait performance, an extensive six- to eight-generation backcross breeding program is needed to introduce the trait into the elite variety, a time-consuming process.
At Syngenta, a new haploid induction editing technology called HI-Edit™ sidesteps the challenge of elite transformation (Kelliher et al. Nat. Biotechnol. 2019; 37: 287–292). HI-Edit introduces the CRISPR-Cas9 machinery into a pollen-generating, easily transformed plant and then crosses that pollen to a recipient elite variety.
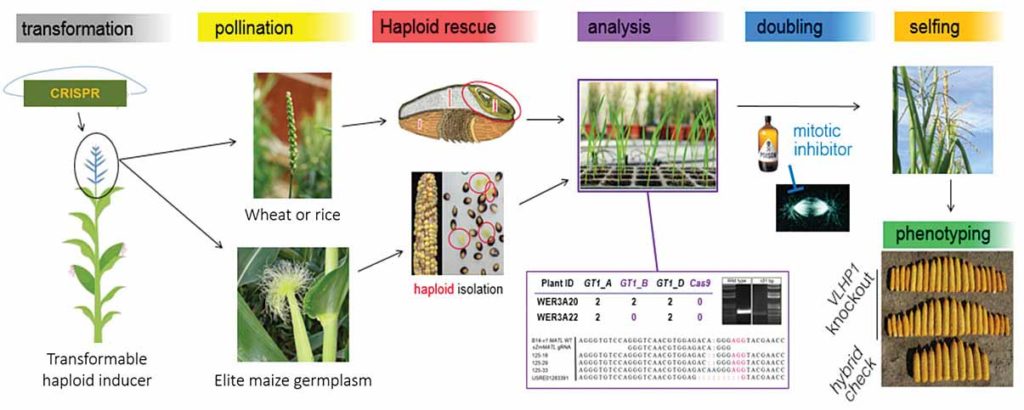
Haploid induction is a modified breeding strategy developed in corn where, in strains called “inducers,” pollen is unsuccessful in fertilizing the egg of a female plant, yet the embryo still grows, albeit with only the maternal half of the expected chromosomes (a haploid). HI-Edit can take advantage of this system because, as the CRISPR-Cas9-bearing pollen attempts to fertilize the recipient egg, the CRISPR machinery makes the edit in the female genome. The (now edited) haploid embryo can then grow into a mature haploid plant, which though fragile is viable.
An additional benefit of this process is that, with a simple colchicine treatment, the haploid plant can duplicate its chromosomes to the correct number. But since the duplicates are all also from the mother, the plant is now homozygous for every gene—including the edited one.
“To introduce a beneficial trait into all of your top varieties in different geographic locations, you could use HI-Edit, a one-step, pollen-delivery system, to make a single cross, and then you would have that trait in an elite background with no transformation process and no gene editing of the recipient line,” explains Ian Jepson, PhD, head of trait research and developmental biology and head of RTP site business at Syngenta.
HI-Edit has been demonstrated in both corn and wheat. In wheat, pollen from an inducer corn strain is crossed to wheat. The pollen then starts to germinate and attempts to fertilize the wheat egg. Although fertilization fails, the CRISPR machinery is delivered via the pollen grain and edits the egg’s nuclear DNA. From a single transformation in corn, Syngenta can now edit wheat without a complicated breeding process.
“Using a different haploid induction system and a gene called CENH3, we have demonstrated the system in Arabidopsis and are looking to exemplify the technology in broad leaf and other crops,” Jepson adds. “The ability to edit crops has been around for a couple of decades, but now we have more efficient, cost-effective tools. As a discovery tool, CRISPR can be used to systematically knock out or modify genes to determine their functions, for example, in the fruit-ripening process.”
Single-molecule sensors
Improvements to plant quality and desirability don’t just come through gene editing. Electrical single-molecule sensors can be applied in the field to increase crop yield and improve plant health. Such sensors are available from Ontera. By applying solid-state nanopore technology, the company ensures that its sensors are cost effective and robust. They can be used in rugged environments to identify seed traits, single nucleotide polymorphisms, soil pathogens, and more.
“If there is something wrong with your car, you do not want the mechanic running every diagnostic test, says William Dunbar, PhD, co-founder, chief technology officer, Ontera. “It would take too long and cost too much. You want a targeted fast analysis.”
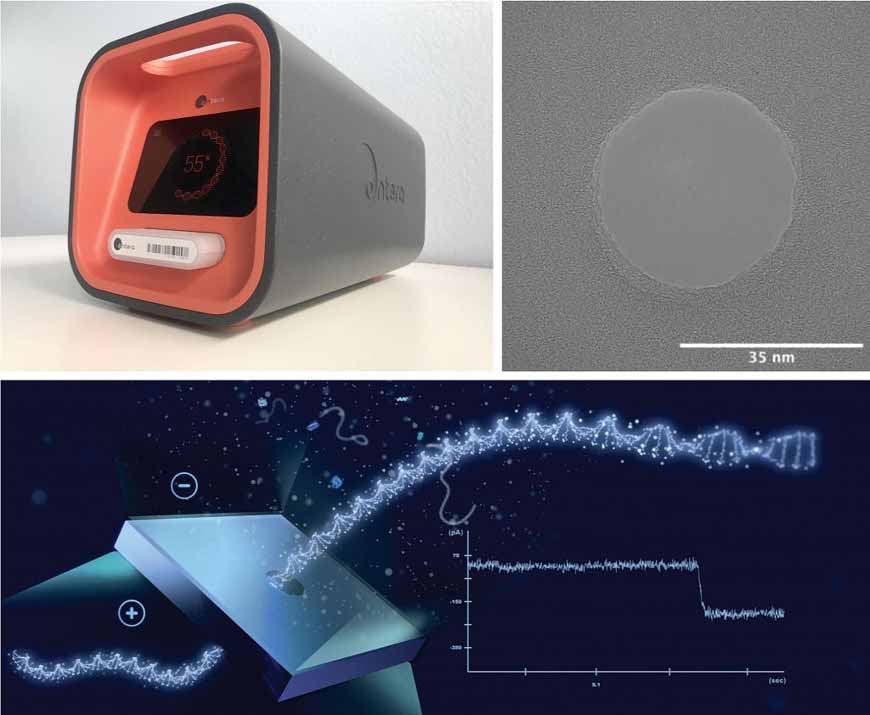

The company has developed a fully integrated sample-to-answer platform called SAM (Sample prep, Amplification, Measure). It runs multiplexed hypothesis-based assays that can detect molecules from 50–50,000 base pairs in 10–15 minutes.
To start the process, a crude sample source is directly inserted into a consumable cartridge. Then the cartridge is inserted into the SAM instrument, which performs endpoint PCR amplification, after which the molecular material is moved from one microfluidic volume to another through a nanopore in a silicon chip. Because a voltage is applied across the nanopore, single molecules are sucked in and pass through, briefly disrupting the current. The duration of the disruption indicates molecular size. Additional information may be gleaned from sample molecules that bind with engineered payload molecules before passing through the nanopore.
“Our focus is bringing lab-quality instrumentation into the field so people can test for molecular ground truth,” says Dunbar. “You need a precise diagnosis to apply a precise treatment.”
The SAM laboratory version will be released in 2020 and made available to developmental partners in 2021. The main agricultural applications are pathogen identification, pathogen resistance, and trait identification—mutations, genetic modifications, or single nucleotide polymorphisms.
Another instrument under development is DUO Nano, a two-pore RUO platform for epigenetics investigation to distinguish phenotypes with identical genome sequences—including base methylation and, more interesting, histone modification.
According to Dunbar, the problem of sequencing is that you know the four-letter options but not the order. Similarly, the order of the modification pattern of histones within a nucleosome string is unknown, but there is a shortlist of key candidates.
A different copy of the same genomic region from a different cell can have a different histone modification profile. Therefore, interrogating each molecule thoroughly is important; DUO Nano will read the same molecule hundreds of times where necessary. Ontera believes that its technology is part of a wave of new technologies that is taking molecular sensing, to use a photography analogy, from shooting on film to creating digital files.
Source: Genetic Engineering and Biotechnology News
By MaryAnn Labant, GEN
